Heart failure (HF) is a clinical syndrome characterized by shortness of breath, fatigue, and fluid retention wherein cardiac output is unable to meet the metabolic needs of the body.1 Despite advances in prevention and management of cardiovascular diseases (CVD), the global prevalence of HF continues to rise due to an increase in comorbidities such as diabetes mellitus (DM), hypertension (HT), and aging populations.2 Understanding the pathophysiology of HF is essential for formulating individualized treatment strategies and reducing cardiovascular risk.3
Growing evidence indicates that inflammatory and autoimmune mechanisms contribute to the etiology and development of several cardiovascular conditions.4,5 Autoimmune diseases can affect single organs or multiple systems.6 Associations between HF and autoimmune diseases, including systemic lupus erythematosus (SLE), ankylosing spondylitis, Crohn’s disease, and celiac disease (CD), have been reported.7
Human leukocyte antigens (HLA)-DQ2 and HLA-DQ8 are well-known genetic risk factors for autoimmune diseases.8 However, their possible roles in other diseases and presence in healthy individuals have not been fully investigated.8 HLA-DQ2 and HLA-DQ8 are useful diagnostic markers for CD and reduce the need for duodenal biopsy.9 A Turkish study reported potential deterioration of cardiac functions in children diagnosed with CD.10 A recent prospective study from Sweden associated HLA-DQ2 and HLA-DQ8 positivity with early vascular atherosclerosis in children with type 1 diabetes under inflammatory conditions.11
Dilated cardiomyopathy has traditionally been attributed to viral infections in many cases, but evidence from animal model studies suggests that autoimmune mechanisms associated with HLA-DQ alleles may also play a role.12 These findings should be interpreted as hypothesis-generating, as direct evidence in human populations remains limited. HLA alleles are not only associated with CD, but also with important cardiovascular problems such as HF, arrhythmia, and atherosclerosis. Thus, it is possible that HLA-DQ2 and HLA-DQ8 haplotypes may be involved in autoimmune susceptibility, which could have implications for CVD development.
These haplotypes’ involvement in CVD with autoimmune or inflammatory components, such as dilated cardiomyopathy and myocarditis, has been increasingly explored in experimental animal models. For example, such a study has demonstrated associations between certain HLA genotypes (including HLA-DQ) and autoimmune responses in dilated cardiomyopathy, as well as a predisposition to immune-mediated myocardial damage.13 Similarly, another study on mice has highlighted the role of HLA-associated immune mechanisms in myocarditis, linking these haplotypes to inflammatory pathways and clinical outcomes.14 The findings from these animal model studies suggest that HLA-DQ2 and HLA-DQ8 may contribute to the pathogenesis of HF via immune dysregulation and warrant further investigation in human HF populations.
However, the presence and role of HLA in HF patients have not been adequately studied. In this study we investigate the relationship between HLA-DQ2 and HLA-DQ8 and HF with reduced ejection fraction (HFrEF).
Methods
This retrospective case–control study was conducted at a tertiary healthcare center between August 2021 and August 2022. The study included 50 patients with HFrEF, defined as a left ventricular ejection fraction (LVEF) < 40%. The patient group was classified into two subgroups, ischemic and non-ischemic, based on coronary angiographic findings and prior coronary revascularization. Controls comprised 50 age- and sex-matched apparently healthy individuals with an LVEF ≥ 40% and no overt or subclinical CVD.
Exclusion criteria for the patient group were the following: (1) age < 18 years or unwillingness to participate; (2) acute or chronic liver disease, severe renal failure defined as estimated glomerular filtration rate (eGFR) of <30 mL/min/1.73 m2); (3) hepatitis B or C infection, inflammatory and hematological diseases, active thyroid disease, cancer, autoimmune thyroiditis, collagen tissue diseases; (4) heart valve disease (moderate or severe), genetic cardiomyopathy (hypertrophic cardiomyopathy, left ventricular noncompaction, restrictive cardiomyopathy), structural heart disease, toxin-induced cardiomyopathy, or peripartum cardiomyopathy; (5) suspected pregnancy; and (6) CD.
Patients with CD were specifically excluded due to well-established relationship with HLA-DQ2 and HLA-DQ8. The intention was to avoid confounding from a known HLA-related autoimmune condition, as we were investigating less well-defined associations between HLA-DQ haplotypes and HF.
Ethical approval was obtained from the Clinical Research Ethics Committee at Çanakkale Onsekiz Mart University, Çanakkale, Turkey (Ref. 2021 08 dated 3/11/2021). The study adhered to the Declaration of Helsinki. Written informed consent was obtained from all participants.
All patients and control participants underwent echocardiographic examinations using a Vivid 7 Pro device (GE, Vingmed, Horten, Norway). Two experienced cardiologists blinded to the study assessed LVEF, which was calculated using the modified Simpson method.15
Samples of blood were taken from peripheral veins via venipuncture and placed in EDTA tubes. Following DNA extraction from peripheral blood using QIAamp DNA Blood Mini Kit, rea- time polymerase chain reaction (PCR) was performed with geneMAP Celiac detection kit and Qiagen Rotor-Gene Q protocol. After the completion of the run, the data were analyzed using melting curve analysis with Rotor-Gene Q series software (QIAGEN) to detect HLA haplotypes.
Statistical analyses were performed using IBM SPSS Statistics for Windows (IBM Corp. Released 2010. IBM SPSS Statistics for Windows, Version 19.0. Armonk, NY: IBM Corp.). The distribution of continuous variables was analyzed using Kolmogorov–Smirnov test. Data were expressed as mean ± SD. Parameters showing normal distribution were compared using the Student’s t-test. Categorical variables were reported as percentages and numbers. Chi-square or Fisher’s exact tests were used to compare the probability ratios of categorical variables. A p-value of < 0.05 was considered statistically significant.
Results
The study included a total of 100 adults: 50 patients with HFrEF (36 men, 14 women) and 50 controls (28 men, 22 women). The mean age of the patient group was 62.2 ± 15.1 years, and that of the control group was 59.7 ± 13.6 years. The patient and control groups had 8 (16.0%) and 10 (20.0%) HLA-DQ2–positive patients, respectively, and 12 (24.0%) and 13 (26.0%) patients with HLA-DQ8–positive patients. Intergroup differences were not significant (p > 0.050). HFrEF patients were classified into two subgroups: ischemic (n = 27) and non-ischemic (n = 23). The ischemic group consisted of patients who had previously undergone percutaneous coronary intervention (n = 18) or coronary artery bypass surgery (n = 9) [Table 1].
Table 1: Demographic and clinical parameters of participants.
|
Demographic
|
|
Age (years)
|
62.2 ± 15.0
|
59.6 ± 13.6
|
0.375
|
|
BMI (kg/m2)
|
25.3 ± 3.8
|
25.2 ± 1.4
|
0.810
|
|
Smoking n (%)
|
8 (16.0)
|
-
|
|
|
Sex
|
|
|
0.145
|
|
Male
|
36
|
28
|
|
|
Female
|
14
|
22
|
|
|
Comorbidities
|
|
Diabetes mellitus n (%)
|
8 (16.0)
|
-
|
|
|
Hypertension n (%)
|
9 (18.0)
|
-
|
|
|
Cardiac parameters
|
|
LVEF (%)
|
28.3 ± 7.9
|
57.6 ± 5.4
|
< 0.001*
|
|
Heart rate (beats/min)
|
83.4 ± 12.1
|
81.2 ± 11.8
|
0.362
|
|
SBP (mmHg)
|
119.4 ± 20.5
|
125.6 ± 12.9
|
0.076
|
|
DBP (mmHg)
|
72.4 ± 8.1
|
72.7 ± 8.3
|
0.837
|
|
Heart failure etiology (HFrEF only)
|
|
Ischemic total
|
27 (54.0)
|
|
|
|
Ischemic (Percutaneous coronary intervention) n (%)
|
18 (36.0)
|
|
|
|
Ischemic (Coronary artery bypass graft surgery) n (%)
|
9 (18.0)
|
|
|
|
Non-ischemic n (%)
|
23 (46.0)
|
|
|
|
HLA status
|
|
HLA-DQ2 n (%)
|
8 (16.0)
|
10 (20.0)
|
0.795
|
HFrEF: heart failure with reduced ejection fraction; LVEF: left ventricular ejection fraction; BMI: body mass index; SBP: systolic blood pressure; DBP: diastolic blood pressure; HLA: human leukocyte antigens.
*This apparent significance is not valid, being due to preselection.
Representative real-time PCR melting curves as positive or negative for HLA-DQ2 and HLA-DQ8 are shown in Figure 1. Each panel represents an individual patient.
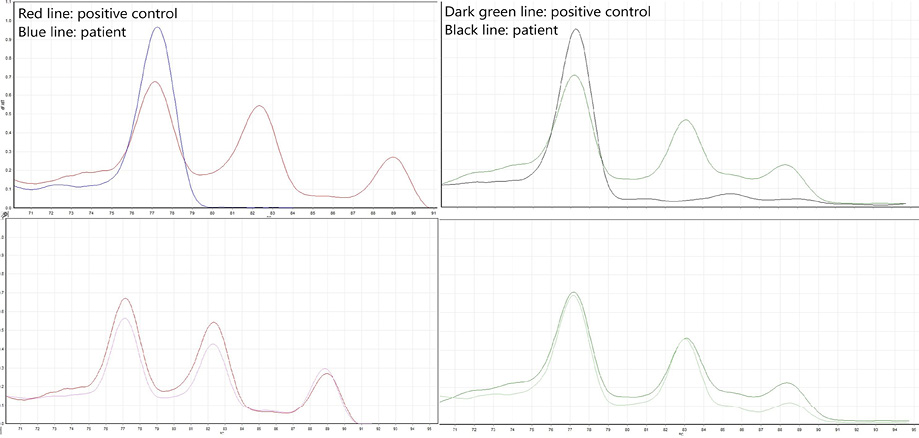 Figure 1: Representative real-time polymerase chain reaction melting curves for HLA-DQ2 and HLA-DQ8 genotyping, showing individual sample traces compared with assay positive controls to determine haplotype positivity or negativity. (a) HLA-DQ2 negative sample showing a single peak not corresponding to the positive control. (b) HLA-DQ2 positive sample showing a characteristic three-peak pattern matching the positive control. (c) HLA-DQ8 negative sample with a single non-matching peak. (d) HLA-DQ8 positive sample demonstrating a three-peak pattern consistent with the positive control.
Figure 1: Representative real-time polymerase chain reaction melting curves for HLA-DQ2 and HLA-DQ8 genotyping, showing individual sample traces compared with assay positive controls to determine haplotype positivity or negativity. (a) HLA-DQ2 negative sample showing a single peak not corresponding to the positive control. (b) HLA-DQ2 positive sample showing a characteristic three-peak pattern matching the positive control. (c) HLA-DQ8 negative sample with a single non-matching peak. (d) HLA-DQ8 positive sample demonstrating a three-peak pattern consistent with the positive control.
Table 2 presents the sex-stratified prevalence of HLA-DQ2 and HLA-DQ8 haplogroups in patients with HFrEF and in controls. HLA-DQ2 positivity was observed in two female and six male in the HFrEF patients and in four women and six men in the control group. HLA-DQ8 was positive in four female and eight male patients in the HFrEF group and in four female and nine male patients in the control group, with no significant differences between groups. Additionally, no significant differences was observed between ischemic and non-ischemic subgroups [Table 2].
Table 2: Sex-stratified prevalence of HLA-DQ2 and HLA-DQ8 haplogroups in HFrEF patients and controls.
|
HFrEF patients (n = 50)
|
|
Male
|
6
|
30
|
> 0.990
|
8
|
28
|
0.718
|
|
Female
|
2
|
12
|
> 0.990
|
4
|
10
|
0.339
|
|
HFrEF patients (ischemic) (n = 27)
|
|
Male
|
4
|
19
|
> 0.990
|
4
|
19
|
> 0.990
|
|
Female
|
0
|
4
|
> 0.990
|
1
|
3
|
> 0.990
|
|
HFrEF patients (non-ischemic) (n= 23)
|
|
Male
|
2
|
11
|
> 0.990
|
4
|
9
|
> 0.990
|
|
Female
|
2
|
8
|
> 0.990
|
3
|
7
|
> 0.990
|
|
Control group (n = 50)
|
|
Male
|
6
|
22
|
> 0.990
|
9
|
19
|
0.339
|
HFrEF: heart failure with reduced ejection fraction; HLA: human leukocyte antigen.
Discussion
Our study revealed that HLA-DQ2 and HLA-DQ8 haplotype frequencies may be similar in this population of HFrEF patients and healthy controls, regardless of sex. As similar trend emerged between ischemic and non-ischemic HFrEF patients in terms of HLA-DQ2 and HLA-DQ8 haplotype frequencies. These findings suggest that HLA haplotypes do not play a role in HFrEF patients, and HLA cannot be used alone as a sufficient marker for early detection of HFrEF.
HLA-DQ2 and HLA-DQ8 haplotypes have been well-recognized by various studies for their association with autoimmune diseases, particularly CD.16 However, their prevalence and diagnostic relevance vary between populations, which may limit their broader applicability.17,18 In children with CD, the aortic strain and stiffness parameters have been found to be similar to those of healthy individuals.19 A Belgian study suggested that HLA-DQA1(a haplotype not considered in our study) may be a genetic risk factor for abdominal aortic aneurysm (AAA).20 A large Swedish study, > 29 000 patients, found that the risk of idiopathic dilated HF potentially increased in patients with CD, albeit slightly below statistical significance (p = 0.052).21
Although our study did not directly investigate the mechanisms by which HLA-DQ2 and HLA-DQ8 may influence cardiovascular outcomes, existing literature suggests potential pathways involving immune-mediated processes. HLA haplotypes, including DQ2 and DQ8, have been associated with autoimmune diseases, which may modulate antigen presentation and T-cell activation.22 Such mechanisms could theoretically contribute to chronic inflammation or autoimmune myocarditis, both of which are implicated in HF pathophysiology. For example, studies have reported associations between HLA-DQ haplotypes and altered cytokine profiles, including increased T-cell stimulation and interleukin-6, which are known to exacerbate myocardial remodeling and dysfunction.23,24 Furthermore, these haplotypes may predispose individuals to immune dysregulation, potentially triggering pathways that impair cardiac function. Future studies should explore these mechanisms through larger cohorts and experimental models to determine how HLA-DQ2 and HLA-DQ8 haplotypes may contribute to a better understanding of the development and progression of HFrEF. Investigating their role in inflammatory and autoimmune processes could provide insights into personalized therapeutic strategies.
This study has several limitations. First, it was conducted at a single center in Western Türkiye, limiting the generalizability of its findings. Second, the small sample size may have reduced the statistical power to detect subtle genetic associations. Third, analysis was limited to HLA-DQ2 and HLA-DQ8 haplotypes; other HLA alleles, such as HLA-DQA1, were not assessed. Fifth, although patients with known CD were excluded, none of the participants underwent a biopsy for CD; therefore, presence of CD patients in the sample cannot be ruled out. Sixth, when comparing between the subgroups of ischemic and non-ischemic HFrEF, finer statistical adjustments such as complex regression analyses or multivariate methods were not performed. Finally, cardiac magnetic resonance imaging was not performed in all HFrEF patients, it was performed only in those with suspected infiltrative disease.
Future studies should build on the present findings by including larger, multicenter cohorts, and more diverse populations, as well as by expanding genetic analyses beyond selected HLA haplotypes. Furthermore, genome-wide association studies could offer valuable insights into other genetic variants that contribute to HF, helping to validate and expand upon our findings.25 Integrating genetic data with functional and immunological analyses may further improve understanding of the mechanisms underlying HF and support the development of more personalized risk stratification and therapeutic strategies.
Conclusion
In this first case–control study of patients from Western Türkiye with HFrEF, no significant association was observed between HFrEF and the presence of HLA-DQ2 or HLA-DQ8 haplotypes, indicating that these variants are unlikely to serve as standalone genetic markers for HFrEF in this population. This negative finding is informative in itself, underscoring the complex and heterogeneous nature of HF, in which individual candidate genes may have limited predictive value. As the genetic influence in the mechanism of HF continues to be unraveled, future studies should move beyond narrow candidate-gene approaches towards larger multi-center designs, incorporating additional genetic markers and their interactions with environmental and clinical modifiers. Such efforts will be essential to clarify the broader implications of genetic markers in the risk stratification, diagnosis, and management of HF in different demographics and generating personalized management strategies.
Disclosure
The authors declare no conflicts of interest. This study was funded by the Scientific Research Coordination Unit of Çanakkale Onsekiz Mart University (Project No. TSA-2022-3876). The sponsors had no role in the study design, data collection, analysis, interpretation, manuscript preparation, or the decision to submit the manuscript for publication.
references
- 1. Heidenreich PA, Bozkurt B, Aguilar D, Allen LA, Byun JJ, Colvin MM, et al. 2022 AHA/ACC/HFSA guideline for the management of heart failure: a report of the American college of cardiology/American heart association joint committee on clinical practice guidelines. Circulation 2022;145(18):e895-e1032.
- 2. Savarese G, Lund LH. Global public health burden of heart failure. Card Fail Rev 2017 Apr;3(1):7-11.
- 3. Schwinger RH. Pathophysiology of heart failure. Cardiovasc Diagn Ther 2021 Feb;11(1):263-276.
- 4. Simmonds SJ, Cuijpers I, Heymans S, Jones EA. Cellular and molecular differences between HFpEF and HFrEF: a step ahead in an improved pathological understanding. Cells 2020 Jan;9(1):242.
- 5. Spahic A, Chen TH, Geller JC, Saenger J, Ohlow MA. Life in the fast lane: clinical and immunohistological characteristics of tachycardia-induced cardiomyopathy-a retrospective study in 684 patients. Herzschrittmacherther Elektrophysiol 2020 Sep;31(3):292-300.
- 6. Zubkiewicz-Kucharska A, Jamer T, Chrzanowska J, Akutko K, Pytrus T, Stawarski A, et al. Prevalence of haplotype DQ2/DQ8 and celiac disease in children with type 1 diabetes. Diabetol Metab Syndr 2022 Sep;14(1):128.
- 7. Kim CH, Tofovic D, Chami T, Al-Kindi SG, Oliveira GH. Subtypes of heart failure in autoimmune diseases. Journal of Cardiac Failure 2017;23(8):S22.
- 8. Kårhus LL, Thuesen BH, Skaaby T, Rumessen JJ, Linneberg A. The distribution of HLA DQ2 and DQ8 haplotypes and their association with health indicators in a general Danish population. United European Gastroenterol J 2018 Jul;6(6):866-878.
- 9. Aboulaghras S, Piancatelli D, Taghzouti K, Balahbib A, Alshahrani MM, Al Awadh AA, et al. Meta-analysis and systematic review of HLA DQ2/DQ8 in adults with celiac disease. Int J Mol Sci 2023 Jan;24(2):1188.
- 10. Erolu E, Polat E. Postural orthostatic tachycardia syndrome in pediatric patients with celiac disease and relationship with tissue transglutaminase antibody levels and HLA tissue group. Haydarpaşa Numune Medical Journal 2020;60(3):211-214.
- 11. Odermarsky M, Pesonen E, Sorsa T, Lernmark Å, Pussinen PJ, Liuba P. HLA, infections and inflammation in early stages of atherosclerosis in children with type 1 diabetes. Acta Diabetol 2018 Jan;55(1):41-47.
- 12. Boldizsar F, Tarjanyi O, Olasz K, Hegyi A, Mikecz K, Glant TT, et al. FTY720 (Gilenya) treatment prevents spontaneous autoimmune myocarditis and dilated cardiomyopathy in transgenic HLA-DQ8-BALB/c mice. Cardiovasc Pathol 2016;25(5):353-361.
- 13. Taneja V, David CS. Spontaneous autoimmune myocarditis and cardiomyopathy in HLA-DQ8.NODAbo transgenic mice. J Autoimmun 2009;33(3-4):260-269.
- 14. Taylor JA, Havari E, McInerney MF, Bronson R, Wucherpfennig KW, Lipes MA. A spontaneous model for autoimmune myocarditis using the human MHC molecule HLA-DQ8. J Immunol 2004 Feb;172(4):2651-2658.
- 15. Lang RM, Badano LP, Mor-Avi V, Afilalo J, Armstrong A, Ernande L, et al. Recommendations for cardiac chamber quantification by echocardiography in adults: an update from the American society of echocardiography and the European association of cardiovascular imaging. Eur Heart J Cardiovasc Imaging 2015 Mar;16(3):233-270.
- 16. Megiorni F, Mora B, Bonamico M, Barbato M, Nenna R, Maiella G, et al. HLA-DQ and risk gradient for celiac disease. Hum Immunol 2009 Jan;70(1):55-59.
- 17. Alarida K, Harown J, Di Pierro MR, Drago S, Catassi C. HLA-DQ2 and -DQ8 genotypes in celiac and healthy Libyan children. Digestive and Liver Disease 2010;42(6):425-427.
- 18. Castro-Antunes MM, Crovella S, Brandão LA, Guimaraes RL, Motta ME, Silva GA. Frequency distribution of HLA DQ2 and DQ8 in celiac patients and first-degree relatives in Recife, northeastern Brazil. Clinics (Sao Paulo) 2011;66(2):227-231.
- 19. Alkan F, Dogan G, Kasırga E, Coskun S. The effect of Celiac disease on cardiac functions and aortic elasticity parameters in children. Cardiol Young 2021 Apr;31(4):627-630.
- 20. Ogata T, Gregoire L, Goddard KA, Skunca M, Tromp G, Lancaster WD, et al. Evidence for association between the HLA-DQA locus and abdominal aortic aneurysms in the Belgian population: a case control study. BMC Med Genet 2006 Jul;7:67.
- 21. Emilsson L, Andersson B, Elfström P, Green PH, Ludvigsson JF. Risk of idiopathic dilated cardiomyopathy in 29 000 patients with celiac disease. J Am Heart Assoc 2012 Jun;1(3):e001594.
- 22. Lerner A, Benzvi C, Vojdani A. HLA-DQ2/8 and COVID-19 in celiac disease: boon or bane. Microorganisms 2023 Dec;11(12):2977.
- 23. Vader W, Stepniak D, Kooy Y, Mearin L, Thompson A, van Rood JJ, et al. The HLA-DQ2 gene dose effect in celiac disease is directly related to the magnitude and breadth of gluten-specific T cell responses. Proc Natl Acad Sci U S A 2003 Oct;100(21):12390-12395.
- 24. Wang J, Wang M, Lu X, Zhang Y, Zeng S, Pan X, et al. IL-6 inhibitors effectively reverse post-infarction cardiac injury and ischemic myocardial remodeling via the TGF-β1/Smad3 signaling pathway. Exp Ther Med 2022 Jul;24(3):576.
- 25. Shah S, Henry A, Roselli C, Lin H, Sveinbjörnsson G, Fatemifar G, et al. Genome-wide association and Mendelian randomisation analysis provide insights into the pathogenesis of heart failure. Nat Commun 2020 Jan;11(1):163.